

Journal für Kulturpflanzen, 74 (11-12). S. 263–270, 2022 | DOI: 10.5073/JfK.2022.11-12.06 | von Maydell et al.
On the search for the vernalization locus in caraway (Carum carvi) using genotyping by sequencing data
Auf der Suche nach dem Vernalisations-Locus in Kümmel (Carum carvi) mittels Genotyping by Sequencing
 | (c) The author(s) 2022 This is an Open Access article distributed under the terms of the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/deed.en). |
Submitted/accepted for publication: 1 August 2022/4 November 2022 |
Caraway (Carum carvi) can be divided into biennial and annual flowering types. Biennials require a cold stimulus for initiation of flowering, whereas annuals lack any vernalization requirement. Cultivation of annuals is considered advantageous due to better integration into crop rotation and reduced production costs. However, available annual varieties lack a sufficient yield potential and essential oil content. Valuable genetic diversity exists within the biennial genepool, which can be transferred to the annual genepool by crossbreeding. According to literature, F1 plants from crosses between annual and biennial flowering types uniformly show annual flowering. Resulting F2 populations show a Mendelian 3:1 ratio between annual and biennial flowering types. This indicates a monogenic inheritance of vernalization requirement in caraway with dominant annual flowering.
After crossbreeding of annual and biennial flowering types, the recessive allele for biennial flowering can remain within the breeding material over multiple generations. Thus, a genetic marker associated with the vernalization locus would be beneficial to detect the recessive allele in one selection step. To identify markers associated with vernalization requirement, we used genotyping by sequencing (GBS) data of 70 biennial and 67 annual flowering types. We conducted case-control studies and population differentiation-based tests. In total, 60 common single nucleotide polymorphism (SNP) markers were found that were significantly associated with vernalization requirement. Out of these, we developed 19 diagnostic markers. In future, these diagnostic markers might be tested in segregating populations.
Horticultural plant, Apiaceae, Umbelliferae, marker-assisted selection (MAS), reproductive trait, Mendel
Kümmel (Carum carvi) kann in zweijährige und einjährige Blühtypen unterteilt werden. Zweijährige benötigen einen Kältereiz für die Blühinduktion, während Einjährige keinen Vernalisationsbedarf haben. Aufgrund der besseren Integration in die Fruchtfolge und der geringeren Produktionskosten wird der Anbau Einjähriger als vorteilhaft angesehen. Die verfügbaren einjährigen Sorten weisen jedoch ein unzureichendes Ertragspotential und einen zu geringen Gehalt an ätherischem Öl auf. Der zweijährige Genpool enthält wertvolle genetische Diversität, die durch Kreuzungen auf den einjährigen Genpool übertragen werden kann. Laut Literatur zeigen F1-Pflanzen aus Kreuzungen zwischen ein- und zweijährigen Blühtypen ein uniform einjähriges Blühverhalten. Resultierende F2-Populationen zeigen ein Mendel'sches 3:1-Spaltungsverhältnis zwischen einjährigen und zweijährigen Blühtypen. Dies deutet auf eine monogene Vererbung des Vernalisationsbedarfs bei Kümmel mit dominant einjährigem Blühverhalten hin.
Nach Kreuzungen zwischen ein- und zweijährigen Blühtypen kann das rezessive Allel für die zweijährige Blüte über mehrere Generationen im Zuchtmaterial verbleiben. Daher wäre ein genetischer Marker, der mit dem Vernalisations-Locus assoziiert ist, vorteilhaft, um das rezessive Allel in einem Selektionsschritt zu erkennen. Um Marker zu identifizieren, die mit dem Vernalisations-Locus assoziiert sind, wurden Genotyping by Sequencing (GBS)-Daten zu 70 zweijährigen und 67 einjährigen Blühtypen genutzt. Case-Control-Studien und populationsgenetische Studien wurden durchgeführt. Insgesamt wurden 60 gemeinsame Einzelnukleotid-Polymorphismus-Marker (SNPs) gefunden, die signifikant mit dem Vernalisationsbedarf assoziiert waren. Für 19 Marker wurden diagnostische Marker entwickelt. Diese könnten in Zukunft in segregierenden Populationen getestet werden.
Gartenbauliche Kultur, Apiaceae, Umbelliferae, Marker-gestützte Selektion (MAS), reproduktives Merkmal, Mendel
Caraway (Carum carvi L., 2n = 2x = 20, Apiaceae) is an important aromatic plant in European and North-American countries. The species can be divided into biennial and annual flowering types. Biennials require a cold stimulus (vernalization) during winter for initiation of flowering. In order to be receptive for this cold stimulus, biennials have to reach a certain size before winter (Németh, 1998b). Consequently, biennial caraway is sown in spring and harvested in summer of the second year. In contrast, annuals can complete their life circle within one growing season. Annuals are sown in early spring and harvested in autumn. Annual material, which is sufficiently winter-hardy, can also be grown as winter-annual, i.e., sowing in autumn and harvest in summer of the next year. To our best knowledge, only one variety (Aprim, Agritec Ltd.) exists, which combines annual behavior with a sufficient degree of winter-hardiness.
In nature, biennials are widespread across Europe and Asia, whereas annuals are restricted to North Africa and the Near East (Pank, 2012). Biennials have been cultivated in European countries since medieval times, whereas annual varieties were introduced at first in the 1990s. For these annual varieties, an Egyptian origin is assumed (Németh, 1998a). Annual cultivation allows better integration into crop rotation and can potentially reduce production costs. However, grain yield and essential oil content of annuals (the most important quality trait) have to be improved by breeding. For winter-annual growing, improvement in winter-hardiness would be beneficial. Unfortunately, the genetic diversity of annual material is limited. The majority of available caraway germplasm of commercial varieties and genebank accessions consists of biennial flowering types. In future breeding programs, the genetic diversity of the biennial genepool could be used to enrich the annual genepool.
Crossbreeding between annuals and biennials is possible, if synchronization of flowering by artificial vernalization can be achieved. Thus, introgression of genetic material from the biennial to the annual genepool is rather easy. However, resulting progenies have to be selected for annual flowering. Németh (1998b) found nearly complete annual flowering in F1-generation after crossbreeding of annual and biennial flowering types. She discovered a Mendelian ratio of 3:1 in an F2-generation. This indicates a monogenic inheritance of vernalization requirement with dominant annual flowering. During the breeding process, the recessive allele for biennial flowering, i.e. vernalization requirement, can remain over several generations within the breeding material. If a genetic marker for vernalization requirement was found, the recessive allele for biennial flowering could be removed already in the F2-generation.
So far, no diagnostic markers for vernalization requirement or any other traits are available in caraway. For a long period, genetic analysis of caraway was restricted to randomly amplified polymorphic DNA (RAPD) markers (Laribi et al., 2011; Bocianowski & Seidler-Lozykowska, 2012; Seidler-Lozykowska et al., 2014). However, RAPD markers provided only a low marker density and showed a low reproducibility (Grover & Sharma, 2016; Garrido-Cardenas et al., 2018). Due to reduced sequencing costs, next generation sequencing (NGS) methods like genotyping by sequencing (GBS) can be implemented for minor crops. Even if a reference genome is not available as for caraway, a high marker density for genome wide association studies (GWAS) can be achieved. Recently, a GBS was conducted to investigate genetic diversity and population structure in a set of 67 annual and 70 biennial flowering types (von Maydell et al., 2021). In this study, we intend to use these data to identify marker loci, which are associated with vernalization requirement of caraway and to develop diagnostic markers. Therefore, this study will foster the development of diagnostic markers in caraway and will pave the path to the establishment of marker-assisted selection (MAS) approaches in caraway.
The material used in this study was previously described in von Maydell et al. (2021). Briefly, a set of 137 caraway accessions was assembled consisting of 67 annual and 70 biennial flowering types. Annual material mainly consists of inbred lines provided by the JKI (Quedlinburg, Germany) and just a few commercial varieties and genebank accessions. Those annual inbred lines predominately originates from crosses between biennial and annual material. Progenies from these crosses were selected for annual flowering and high essential oil content. In contrast, biennial material mainly consisted of wild material, commercial varieties and genetic resources from genebanks.
Accessions were assigned to a flowering type based on data assessed in a field trial (von Maydell et al., 2021). Accessions that uniformly flowered in the year of sowing were defined as annual flowering types and accessions that uniformly flowered in the year after sowing were defined as biennial flowering types.
Genotyping of the caraway accessions under investigation was previously described in von Maydell et al. (2021). Briefly, GBS was conducted using the method published by Elshire et al. (2011). Lacking a reference genome, a so-called 'mock reference' composed of consensus GBS fragments was constructed (Melo & Hale, 2019). Reads were mapped and a SNP calling was carried out. Finally, SNPs were filtered for quality, ≤ 10% missing values, ≤ 5% minor allele frequency (MAF) and ≤ 90% maximum heterozygosity. This resulted in a marker set of 13,155 SNPs.
To identify loci associated with vernalization requirement, case-control studies based on a Fisher’s exact test were conducted using the software SNPsift (Cingolani et al., 2012). SNPsift allows to test whether the alternative allele is associated with the trait of interest in a dominant or recessive way. However, the alternative allele could predominantly be found either in the annuals or in the biennials. Hence, both, the dominant and the recessive model, are computed for the collection of 137 accessions with 67 annuals (=cases) and 70 biennials (=controls).
Furthermore, population differentiation-based statistics were used to identify genome regions, which indicate shifts in allele frequencies between the group of annuals (67) and biennials (70). For this purpose, Fst-outlier detection tests based on the FDIST algorithm (Beaumont & Nichols, 1996) were conducted by using the software packages Arlequin v.3.5.2.2 (Excoffier & Lischer, 2010). Two different approaches were implemented, i.e., the non-hierarchical finite island model (Arlequin) and the hierarchical island model (Arlequin settings: 20,000 simulations, 100 demes, 10 groups). For the hierarchical island model, a hierarchical structure has to be defined. Therefore, the group of annuals and biennials was randomly divided in two subgroups of 33 and 34 accessions or 35 and 35 accessions, respectively (Liu et al., 2017).
P-values of SNPsift and Arlequin models were adjusted for multiple testing (FDR, Benjamini & Hochberg, 1995) by using the function p.adjust (setting: method = “fdr”) implemented in the software package R (R Core Team, 2019). Markers with FDR adjusted p-values < 0.001 were considered as significant SNPs. Significant markers, which were detected by all four fitted models, were defined as candidate markers to be associated with the vernalization locus.
It was tested whether promising markers detected by SNPsift and Arlequin can be transferred to the PACE marker system (3cr Bioscience). The technique was recently implemented for caraway in von Maydell et al. (2020). Briefly, a PCR with allele specific primers for the SNP and an endpoint fluorescent measurement was performed. Developed markers were tested for functionality by using six biennial samples, eight annual samples and two negative controls. As biennial samples, two plants each of the cultivars Niederdeutscher, Prochan and Rekord were randomly selected. As annual samples, two plants each of the cultivars Aprim, Karzo, Sprinter and SZK-1 were used. DNA was extracted using DNeasy Plant Mini Kit (Qiagen). DNA quantity was estimated using the Qubit 2.0 Fluorimeter (Life Technologies). DNA concentration was set to 10 ng/μl. PCR was carried out as described in von Maydell et al. (2020). The primers of functional PACE markers are listed in Table 1. The corresponding SNP flanking sequences can be extracted from supplementary Table S.
Table 1. List of developed PACE markers for loci significantly associated with vernalization requirement detected by SNPsift and Arlequin models. The locus ID consists of the ID of the contig and the SNP position on the contig. ASP1/ASP2 = allele specific primers, CP = common primer. FAM = oligo sequence GAAGGTGACCAAGTTCATGCT; HEX = oligo sequence GAAGGTCGGAGTCAACGGATT.
Marker ID | Locus ID | Primer type | Oligo sequence |
Cc2021M064 | 18965_39 | ASP1 | FAM-AAGAACAAAGTGTATCACTTCTCAG |
|
| ASP2 | HEX-ACAYAAGAACAAAGTGTATCACTTCTCAA |
|
| CP | GCTATAGGAATTTTGTRTTATTGACGCTTT |
Cc2021M067 | 207597_145 | ASP1 | FAM-TTTAAGCCGTAGTCTTGAATCATCG |
|
| ASP2 | HEX-GTTTTAAGCCGTAGTCTTGAATCATCA |
|
| CP | GCTGGTAGCWGAAGGGACAAGTTTT |
Cc2021M068 | 9780_153 | ASP1 | FAM-AATGGCTTCCTCCTCCTTTGAGA |
|
| ASP2 | HEX-ATGGCTTCCTCCTCCTTTGAGG |
|
| CP | AGAGATTGCCCCCTGTTTTTCTAGATATA |
Cc2021M070 | 478799_86 | ASP1 | FAM-ATTTGTTAGACATTACAGCAACACTCTTT |
|
| ASP2 | HEX-GTTAGACATTACAGCAACACTCTTG |
|
| CP | TGGGCTCATGACCAGCTCAAACTAA |
Cc2021M077 | 245881_125 | ASP1 | FAM-GAAGTGTGCTAACTTTGCTCTGC |
|
| ASP2 | HEX-GTGAAGTGTGCTAACTTTGCTCTGT |
|
| CP | CCAGTGACCAACAGTATAAAGTAGAACAT |
Cc2022M113 | 25780_45 | ASP1 | FAM-CACTTCACTAGTCACAAGACACG |
|
| ASP2 | HEX-GCACTTCACTAGTCACAAGACACA |
|
| CP | GATTTGTGTAATGGCAGACAAAGACAGAA |
Cc2022M117 | 6155_88 | ASP1 | FAM-CGAATACTTTAACGACCAACTTAACGAT |
|
| ASP2 | HEX-GAATACTTTAACGACCAACTTAACGAG |
|
| CP | CTTTTTAACCCATTACGGACGTTTACTTTA |
Cc2022M118 | 7652_114 | ASP1 | FAM-AGAAGCACAGAGCGTGACCAAC |
|
| ASP2 | HEX-GAGAAGCACAGAGCGTGACCAAT |
|
| CP | TTGAGAATAGTTGATCAGGAATAAAGTTCA |
Cc2022M120 | 7652_69 | ASP1 | FAM-TCTCACCTTATGTACTTGTATTTATAT |
|
| ASP2 | HEX-GCTTCTCACCTTATGTACTTGTATTTATAA |
|
| CP | GGTTCACACAAGGAGTCTTTAAACAACTT |
Cc2022M121 | 7652_151 | ASP1 | FAM-TATTCCTGATCAACTATTCTCAAAAACAG |
|
| ASP2 | HEX-CTTTATTCCTGATCAACTATTCTCAAAAACAT |
|
| CP | CACACATCATAATTACTAAATTGCTCTTAT |
Cc2022M123 | 10122_133 | ASP1 | FAM-CTACCTAAGTATCGAAAAAGTAACAG |
|
| ASP2 | HEX-CCTCTACCTAAGTATCGAAAAAGTAACAA |
|
| CP | GGCAATGATAGGAGTTTTCAAGCTGATTT |
Cc2022M124 | 11783_102 | ASP1 | FAM-CGTATCAATCTTCTCCTTATATTCAGGAA |
|
| ASP2 | HEX-GTATCAATCTTCTCCTTATATTCAGGAG |
|
| CP | CTTACTACATCAGCACATGCTAAACTGAT |
Cc2022M125 | 11783_133 | ASP1 | FAM-GTCATCAGTATTTAGCACGTCAGG |
|
| ASP2 | HEX-GTCATCAGTATTTAGCACGTCAGC |
|
| CP | ARATCAGTTTAGCATGTGCTGATGTAGTAA |
Cc2022M126 | 124119_175 | ASP1 | FAM-AATCGTCTCCGGCACTATCCG |
|
| ASP2 | HEX-CAATCGTCTCCGGCACTATCCA |
|
| CP | GTCGGAAAGATTTACGAACATCGTCATAA |
Cc2022M128 | 207597_172 | ASP1 | FAM-GCATCCATAATGCTCAACCTTCA |
|
| ASP2 | HEX-GCATCCATAATGCTCAACCTTCG |
|
| CP | TAAGAYGATGATTCAAGACTACGGCTTAAA |
Cc2022M129 | 214096_140 | ASP1 | FAM-CCTGAATTCTGCTATATGTTTATGCC |
|
| ASP2 | HEX-CCTGAATTCTGCTATATGTTTATGCG |
|
| CP | CTAACATTGCAAAATAACCAAAGAAACATT |
Cc2022M132 | 255309_191 | ASP1 | FAM-ACCTTATAAGGCATTTATCAAAATCATATTA |
|
| ASP2 | HEX-CTACCTTATAAGGCATTTATCAAAATCATATTT |
|
| CP | CGAGTATTAGCTGTGATATATTGCATCAAT |
Cc2022M133 | 261365_69 | ASP1 | FAM-AAGAGCCTTRTAAGGCCACTCAG |
|
| ASP2 | HEX-CAAGAGCCTTRTAAGGCCACTCAT |
|
| CP | AACAACCTTGATCAYAAGAAAGCTCAACAT |
Cc2022M136 | 5784_204 | ASP1 | FAM-CAGGTCATATATTAGACCGCACTC |
|
| ASP2 | HEX-CTCAGGTCATATATTAGACCGCACTA |
|
| CP | GTATTGTGGGTTGAAATCATTGGAGGTTT |
We evaluated the dominant and recessive model used in the case-control study (SNPsift). The dominant and the recessive model revealed 3,411 and 436 markers significantly associated with vernalization requirement, respectively (p value < 0.001). The markers with the lowest p value were locus 7652_114 (dominant model) and locus 9780_153 (recessive model).
In total, 108 and 75 markers were detected as significant SNPs (p value < 0.001) by using the non-hierarchical finite island model or the hierarchical island model in Arlequin, respectively. Identical to the dominant SNPsift model, both models detected locus 7652_114 as peak marker.
In total, 60 significantly associated markers were detected by both Arlequin models and both SNPsift models. This includes the peak markers from all models. These common 60 loci were defined as putative candidates associated with vernalization requirement in caraway. Further information of these loci, including SNP sequences and p values, can be retrieved from supplementary Table S. For several loci, the development of diagnostic markers was tested.
Out of these 60 markers, 19 markers were transferred to the PACE marker system. In a test for functionality with a small set of annual and biennial ecotypes, these markers showed a sufficient allelic discrimination, i.e., at least two distinct clusters could be visually discriminated (three clusters if heterozygous genotypes were present). Exemplarily, an allelic discrimination with three distinct clusters (genotypes) is presented in Figure 1. The primer sequences of these markers are listed in Table 1. Fortunately, marker design was successful for the peak markers of all models. In few cases, allelic discrimination might be further improved by optimizing PCR settings (for instance annealing temperature).
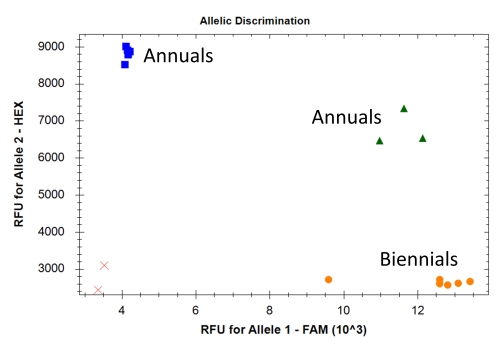
Fig. 1. Exemplary result (allelic discrimination) of the PACE marker Cc2022M132 designed for the SNP locus 255309_191. Red crosses = negative control, blue squares = homozygous allele 2, green triangles = heterozygous genotypes, orange dots = homozygous allele 1. In this case, all biennial samples showed a homozygous allele 1 genotype and annual samples either a homozygous allele 2 genotype or a heterozygous genotype.
For all 19 loci with successful design of diagnostic markers, the allelic distribution based on GBS data within the investigated 67 annuals and 70 biennials is presented in Table 2. All loci showed a distinct but incomplete allelic discrimination between flowering types, including peak markers (Table 2).
Table 2. Allelic distribution of 19 selected SNP markers significantly associated with vernalization requirement detected by SNPsift and Arlequin models. The allelic distribution within the 67 annual accessions and 70 biennial accessions is presented as follows: REF = homozygous reference (first allele in brackets, supplementary Table S) genotypes. ALT = homozygous alternative (second allele in brackets) genotypes. HET = heterozygous genotypes. Missing values are not presented.
Locus ID | Annuals | Biennials | ||||
REF | ALT | HET | REF | ALT | HET | |
7652_114 | 4 | 48 | 13 | 65 | 0 | 1 |
9780_153 | 2 | 50 | 14 | 54 | 0 | 12 |
11783_102 | 6 | 43 | 12 | 66 | 0 | 1 |
11783_133 | 6 | 43 | 12 | 66 | 0 | 1 |
5784_204 | 5 | 37 | 17 | 64 | 0 | 2 |
245881_125 | 5 | 42 | 20 | 68 | 0 | 2 |
214096_140 | 7 | 41 | 16 | 60 | 0 | 5 |
255309_191 | 12 | 43 | 9 | 59 | 0 | 7 |
10122_133 | 7 | 38 | 14 | 62 | 0 | 3 |
207597_145 | 7 | 43 | 14 | 64 | 0 | 5 |
207597_172 | 7 | 43 | 14 | 64 | 0 | 5 |
7652_69 | 4 | 48 | 13 | 63 | 0 | 4 |
7652_151 | 4 | 48 | 13 | 64 | 0 | 2 |
261365_69 | 8 | 33 | 23 | 61 | 0 | 2 |
124119_175 | 4 | 46 | 16 | 52 | 0 | 14 |
478799_86 | 51 | 1 | 14 | 4 | 43 | 14 |
18965_39 | 54 | 1 | 12 | 0 | 47 | 21 |
6155_88 | 61 | 0 | 6 | 3 | 14 | 51 |
25780_45 | 8 | 36 | 23 | 64 | 0 | 3 |
According to literature, vernalization requirement shows a monogenic inheritance with dominant annual flowering (Németh, 1998b). Thus, vernalization requirement in caraway can be regarded as a “simple” Mendelian trait. Since the discovery of the rules of inheritance by Gregor Mendel (Mendel, 1866), further scientific innovations led to the mapping and cloning of many loci encoding the expression of Mendelian traits. However, so far, it was not possible to map a trait-associated locus or to identify a corresponding gene in caraway. Genetic mapping of the vernalization requirement and the development of diagnostic markers could be a good starting point for genetic analysis of traits in caraway.
In a previous study, it could be shown that the vernalization requirement is the main factor that causes population differentiation in caraway (von Maydell et al., 2021). Therefore, it can be assumed that markers associated with the vernalization locus have the strongest effect on population differentiation. Other factors, which are known to affect population differentiation, i.e., breeding history or geographical origin, cause only minor effects in the caraway population under investigation. Furthermore, it can be assumed that markers associated with the vernalization requirement cause the strongest shifts in allele frequencies between the group of annual and biennial caraway accessions. Based on these assumptions, SNPsift and Arlequin models were fitted. These models commonly identified 60 putative candidate loci associated with vernalization requirement in caraway.
Nevertheless, loci associated with vernalization requirement might not necessarily be physically linked with the vernalization locus. However, above 67% of the examined annual accessions originate from crossbreeding between annual and biennial material, which progeny were selected for annual flowering in breeding programs. Due to these rather recent recombination events, we expected that at least markers strongly associated with vernalization requirement should indeed be physically linked to the putative vernalization locus.
Assuming a monogenic inheritance of vernalization requirement in caraway, all candidate markers should be located within the same genomic region. Unfortunately, the position of the markers on the genome is unknown, as a reference genome of caraway is not yet available. Hence, without a reference genome or any genetic map, the location of the candidate markers cannot be determined. Consequently, these markers should be tested in a mapping population to estimate the actual genetic linkage of the markers with the putative vernalization locus. Moreover, this mapping population should be used to design a first genetic map for caraway across all chromosomes. For this purpose, again GBS might be the method of choice to genotype the mapping population. The designed PACE markers can be additionally used for genotyping.
These markers might not yet be in very strong linkage with the vernalization locus, because the detected markers did not perfectly distinguish between annuals and biennials. All markers showed some degree of heterozygosity within one or both groups of flowering types (Table 2). Due to the dominant inheritance of annual flowering, heterozygous plants within annual accessions would be possible. However, in field trials, we evaluated up to 75 plants per accession and did not observe segregating accessions regarding vernalization requirement (von Maydell et al., 2021). Thus, we would not expect heterozygous genotypes for loci closely linked to the vernalization locus even for annual flowering types. Nevertheless, the marker closest to the putative vernalization locus combined with at least one flanking marker on the other side of the vernalization locus could be a good starting point for MAS. Furthermore, the PACE markers could be useful to evaluate if crossbreeding between annual and biennial flowering material was successful. In the future, marker saturation in the genomic region of interest could be conducted to get additional markers closer linked to the vernalization locus (fine mapping).
Finally, it should be of interest to identify the gene that controls vernalization requirement in caraway. In Arabidopsis thaliana, the flowering locus C (FLC) was identified to control vernalization requirement (Michaels & Amasino, 1999). Since then, several other players acting within the vernalization pathway or network have been identified. Several of these players encode MADS-box transcription factors (Whittaker & Dean, 2017). Henceforth, Arabidopsis sequences were used to detect homologs in other species. In carrot, as distant relative of caraway, two different vernalization loci are assumed (Wohlfeiler et al., 2019) and one putative locus has been mapped (Alessandro et al., 2013). In addition, several homologs of players in the vernalization pathway were found (Ou et al., 2017). Due to genome reduction and due to the short contigs of the ‘mock reference’, available GBS data are scarcely suitable to search for homologs. The generation of a reference genome of caraway would considerably simplify such analyses.
Several promising markers putatively associated with vernalization requirement in caraway were identified and transferred to a diagnostic marker system. The applicability of these markers for MAS has to be tested in mapping populations. In general, genetic analysis in caraway is hampered by the lack of a genetic map and a reference genome. Thus, the design of a first genetic map and implementation of MAS would be a milestone for caraway breeding, which should be endeavored in the near future.
We thank Jenny Knibbiche for establishing and phenotyping the field trials and Beate Kamm for the laborious marker tests. We thank all germplasm providers, which enabled the assembly of a diverse caraway collection: The Leibniz Institute of Plant Genetics and Crop Plant Research (IPK, Gatersleben, Germany); the United States Department of Agriculture (USDA, Ames, USA) the Crop Research Institute (CRI, Prague, Czech Republic); the Centre for Genetic Resources, the Netherlands (CGN, Wageningen, Netherlands); the Nordic Genetic Resource Center (NordGen, Alnarp, Sweden); the Julius Kuehn-Institute (JKI, Quedlinburg, Germany); the Dr. Junghanns GmbH (Ascherleben, Germany); the Agritec Ltd. (Šumperk, Czech Republic); the National Research Council Canada (NRC, Ottawa, Canada); the Chrestensen GmbH (Erfurt, Germany); the Pharmasaat GmbH (Artern, Germany).
This project was funded by the "Fachagentur Nachwachsende Rohstoffe" (FNR) on behalf of the Federal Ministry of Food and Agriculture (BMEL) (Funding ID: 22023215).
The authors declare that they do not have any conflicts of interest.
Table S. List of 60 loci significantly associated with vernalization requirement detected by SNPsift and Arlequin models. Contains following information: SNP flanking sequences, reference and alternative allele, minor allele frequency, p values of SNPsift and Arlequin models and rank of p values per model.
The supplementary information (Table S) for this article can be found online at https://doi.org/10.5073/JfK.2022.11-12.06.
Alessandro, M.S., C.R. Galmarini, M. Iorizzo, P.W. Simon, 2013: Molecular mapping of vernalization requirement and fertility restoration genes in carrot. Theoretical and Applied Genetics 126 (2), 415-423, DOI: 10.1007/s00122-012-1989-1.
Beaumont, M.A., R. Nichols, 1996: Evaluating loci for use in the genetic analysis of population structure. Proceedings of the Royal Society of London B: Biological Sciences 263, 1619-1626, DOI: 10.1098/rspb.1996.0237.
Benjamini, Y., Y. Hochberg, 1995: Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society: Series B (Methodological) 57 (1), 289-300, DOI: 10.1111/j.2517-6161.1995.tb02031.x.
Bocianowski, J., K. Seidler-Lozykowska, 2012: The relationship between RAPD markers and quantitative traits of caraway (Carum carvi L.). Ind Crops Prod 36 (1), 135-139, DOI: 10.1016/j.indcrop.2011.08.019.
Cingolani, P., V.M. Patel, M. Coon, T. Nguyen, S.J. Land, D.M. Ruden, X. Lu, 2012: Using Drosophila melanogaster as a Model for Genotoxic Chemical Mutational Studies with a New Program, SnpSift. Front Genet 3, 35, DOI: 10.3389/fgene.2012.00035.
Elshire, R.J., J.C. Glaubitz, Q. Sun, J.A. Poland, K. Kawamoto, E.S. Buckler, S.E. Mitchell, 2011: A Robust, Simple Genotyping-by-Sequencing (GBS) Approach for High Diversity Species. Plos One 6 (5), e19379, 10, DOI: 10.1371/journal.pone.0019379.
Excoffier, L., H.E. Lischer, 2010: Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10 (3), 564-567, DOI: 10.1111/j.1755-0998.2010.02847.x.
Garrido-Cardenas, J.A., C. Mesa-Valle, F. Manzano-Agugliaro, 2018: Trends in plant research using molecular markers. Planta 247 (3), 543-557, DOI: 10.1007/s00425-017-2829-y.
Grover, A., P.C. Sharma, 2016: Development and use of molecular markers: past and present. Critical Reviews in Biotechnology 36 (2), 290-302, DOI: 10.3109/07388551.2014.959891.
Laribi, B., N. Zoghlami, M. Lamine, K. Kouki, A. Ghorbel, A. Mougou, 2011: RAPD-based assessment of genetic diversity among annual caraway (Carum carvi) populations. Eurasian J Biosci 5 (1), 37-47.
Liu, L., K.P. Ang, J.A. Elliott, M.P. Kent, S. Lien, D. MacDonald, E.G. Boulding, 2017: A genome scan for selection signatures comparing farmed Atlantic salmon with two wild populations: Testing colocalization among outlier markers, candidate genes, and quantitative trait loci for production traits. Evol Appl 10 (3), 276-296, DOI: 10.1111/eva.12450.
Melo, A.T.O., I. Hale, 2019: Expanded functionality, increased accuracy, and enhanced speed in the de novo genotyping-by-sequencing pipeline GBS-SNP-CROP. Bioinformatics 35 (10), 1783-1785, DOI: 10.1093/bioinformatics/bty873.
Mendel, G., 1866: Versuche über Pflanzen-Hybriden. Verhandlungen des Naturforschenden Vereines in Brünn. Brünn, Naturforschender Verein, 4, 3-47.
Michaels, S.D., R.M. Amasino, 1999: <em>FLOWERING LOCUS C</em> Encodes a Novel MADS Domain Protein That Acts as a Repressor of Flowering. The Plant Cell 11 (5), 949-956, DOI: 10.1105/tpc.11.5.949.
Németh, É., 1998a: Introduction. In: Nemeth, E. (Ed.), Caraway The Genus Carum. Amsterdam, harwood academic publishers, 7, 1-8.
Németh, É., 1998b: Questions of the generative development. In: Nemeth, E. (Ed.), Caraway The Genus Carum. Amsterdam, harwood academic publishers, 7, 69-81.
Ou, C.G., J.H. Mao, L.J. Liu, C.J. Li, H.F. Ren, Z.W. Zhao, F.Y. Zhuang, 2017: Characterising genes associated with flowering time in carrot (Daucus carota L.) using transcriptome analysis. Plant Biology 19 (2), 286-297, DOI: 10.1111/plb.12519.
Pank, F., 2012: Kümmel (Carum carvi L.). In: Hoppe, B. (Ed.), Handbuch des Arznei- und Gewürzpflanzenanbaus. Bernburg, Verein für Arznei- und Gewürzpflanzen SALUPLANTA e. V., 4, 728-762.
R Core Team, 2019: R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL http://www.R-project.org/.
Seidler-Lozykowska, K., A. Kuczynska, K. Mikolajczyk, J. Nowakowska, J. Bocianowski, 2014: Estimation of genetic distance among genotypes of caraway (Carum carvi L.) using RAPD-PCR. Acta Sci Agron 36 (2), 183-188, DOI: 10.4025/actasciagron.v36i2.18197.
von Maydell, D., J. Brandes, H. Lehnert, W. Junghanns, F. Marthe, 2020: Breeding synthetic varieties in annual caraway: observations on the outcrossing rate in a polycross using a high-throughput genotyping system. Euphytica 217 (1), 1, DOI: 10.1007/s10681-020-02732-5.
von Maydell, D., H. Lehnert, T. Berner, E. Klocke, W. Junghanns, J. Keilwagen, F. Marthe, 2021: On genetic diversity in caraway: Genotyping of a large germplasm collection. PLoS One 15 (12), e0244666, DOI: 10.1371/journal.pone.0244666.
Whittaker, C., C. Dean, 2017: The FLC Locus: A Platform for Discoveries in Epigenetics and Adaptation. Annual Review of Cell and Developmental Biology 33 (1), 555-575, DOI: 10.1146/annurev-cellbio-100616-060546.
Wohlfeiler, J., M.S. Alessandro, P.F. Cavagnaro, C.R. Galmarini, 2019: Multiallelic digenic control of vernalization requirement in carrot (Daucus carota L.). Euphytica 215 (2), 37, DOI: 10.1007/s10681-019-2360-2.