

Journal für Kulturpflanzen, 74 (11-12). S. 223–232, 2022 | DOI: 10.5073/JfK.2022.11-12.02 | Friedt and Ordon
Mendel’s Laws and their impact on plant breeding
Zur Bedeutung der Mendel’schen Regeln für die Pflanzenzüchtung
 | (c) The author(s) 2022 This is an Open Access article distributed under the terms of the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/deed.en). |
Submitted/accepted for publication: 25 August 2022/8 November 2022 |
Cereals like wheat, rice, maize, barley and millets, feed the world. Therefore, global breeding activities, which had been very successful during the last decades, aim at an increase of cereal yields. This, as expected continued success story is the result of the extensive observations and formulation of the fundamental genetic rules that bear his name as Mendel’s law of inheritance (T.H. Morgan 1911). Mendel’s thinking in “heritable characters“ resembling structural “genes“, was the basis for a better understanding of the genetic principles of inheritance; The application of these principles in systematic plant breeding has then allowed the continuous development of improved cultivars.
Plant characteristics controlled by a few or only one gene were the first candidates for improvement since they allowed the direct application of Mendel’s rules. Typical examples are resistances against diseases, e.g. due to fungal pathogens or viruses. Today, most of the wheat and barley cultivars grown in Europe are resistant to many diseases. The discovery of resistance of barley against soil-borne barley yellow mosaic virus disease and the clarification of its genetic control is an impressive example for the direct application of Mendel’s law. The respective extensive research was the basis for developing a multitude of resistant barley varieties during recent decades. Numerous further examples for resistance of crop plants against pathogens could be mentioned, here. Such “Mendel genes" can be genetically marked and localized, which subsequently enables marker-assisted selection. They were also among the first to be isolated. Isolated genes are the basis to apply new breeding technologies, e.g. CRISPR/Cas, and to transfer the respective genes to other varieties, species or taxa with the help of biotechnological tools.
Due to the obviously increasing effects of climate change, it will be necessary in the future to breed new varieties with higher tolerance to abiotic stress – such as heat and drought. Such traits are usually not controlled by one or a few genes; rather, they are polygenically inherited and therefore show a typical quantitative distribution of respective traits. This also applies to crop yield and relevant quality traits. New approaches have been developed for breeding and improving such complex traits e.g. QTL analyses separating complex traits into several Mendelian loci explaining part of the variance observed and „genomic selection" are widely applied today. In this process, suitable genotypes are examined for genetic variation that indicates a desired trait expression (phenotype). In this way, a continuous optimisation of methodology takes place in today's knowledge-based plant breeding on the basis of Mendel's rules via empirical (“classical") methodology. This will be the cornerstone to improve the yield potential, yield stability and quality of plants in the future.
genetics, genotype, haploids, inheritance, Mendel´s law, monogenic traits, phenotype, plant breeding, polygenic traits, polyploidy, quantitative traits
Die landwirtschaftliche Pflanzenproduktion wurde im Laufe des letzten Jahrhunderts erheblich gesteigert; so konnte die weltweite Weizenproduktion seit den 1970er Jahren durch Züchtung und effektivere Produktionstechnik fast verdoppelt werden. Auch die Getreidearten Reis, Mais, Gerste und Hirse haben heute eine große globale Bedeutung als Grundlage für Nahrungs- und Futtermittel.
Diese Erfolgsgeschichte wäre ohne die Erkenntnisse von Gregor Mendel so nicht möglich gewesen. Mendel hat Vererbungsmuster erkannt und beschrieben, die er als „vererbbare Eigenschaften" bezeichnete. Das Denken in Faktoren (d. h. „Genen") war die Grundlage für ein besseres Verständnis der Vererbung von Eigenschaften; Die Anwendung der Mendel‘schen Regeln in der systematischen Pflanzenzüchtung ermöglichte die kontinuierliche Entwicklung neuer Sorten mit verbesserter Resistenz gegen Krankheiten und Schädlinge sowie besserer Produktqualität. Dies war möglich, weil diese Merkmale häufig von wenigen oder auch einzelnen Genen – monogenisch – gesteuert werden. Ein Beispiel für solche „Mendel’schen Gene“ ist die Resistenz der Gerste gegen die bodenbürtige Gelbmosaikvirose. Eine Vielzahl weiterer Beispiele, z. B. Resistenzen gegen Mehltau und Rostkrankheiten, sind bekannt. Diese Gene können mittels molekularer Methoden markiert und lokalisiert und somit in markergestützten Selektionsverfahren genutzt werden. Auch gehörten sie zu den ersten, die physisch isoliert werden konnten. Isolierte Gene sind die Grundlage für die Nutzung neuer Züchtungstechnologien, z. B. CRISPR/Cas, und sie können mithilfe biotechnologischer Verfahren auch auf andere Sorten, Arten oder Taxa übertragen werden.
Aufgrund der offensichtlich zunehmenden Auswirkungen des Klimawandels wird es in Zukunft notwendig sein, neue Sorten mit einer besseren Toleranz gegenüber abiotischem Stress – wie Hitze und Trockenheit – zu züchten. Solche Eigenschaften werden in der Regel nicht von einem oder wenigen Genen kontrolliert; sie sind vielmehr polygenisch vererbt und zeigen daher eine typische quantitative Merkmalsausprägung. Dies gilt auch für die Höhe des Ernteertrags und maßgebliche Qualitätsmerkmale. Für die Züchtung und Verbesserung solch komplexer Merkmale wurden in jüngerer Zeit neue Ansätze entwickelt, so z. B. die QTL Analyse, in der komplexe Merkmale in einzelne Loci zerlegt werden, die einen Teil der beobachteten Varianz erklären; in jüngster Zeit kommen sogenannte genomische Selektionsverfahren hinzu. Auf diese Weise findet in der heutigen wissensbasierten Pflanzenzüchtung auf der Grundlage der Mendel’schen Regeln über die empirische („klassische“) Methodik hinaus eine kontinuierliche Erweiterung und Optimierung des Methodenspektrums statt. Damit wird auch künftig eine Verbesserung des Ertragspotentials, der Ertragsstabilität und der Qualität von pflanzlichen Produkten möglich sein.
Genetik, Genotyp, Haploide, Mendel´sche Regeln, monogenische Merkmale, Phänotyp, Pflanzenzüchtung, polygenische Merkmale, Polyploidie, quantitative Merkmale, Vererbung
Agricultural plant production has increased considerably during the last century. Wheat, rice, maize, barley and millets, have an enormous global importance as the basis of food and feed. For example, wheat yield in central Europe increased from 2 to 8 t/ha in the last century and since the 1970s, world production of wheat has been duplicated and risen in Germany by a factor of 2.3-2.4 (Table 1). This is primarily due to the enhancement of yield by breeding and improved agricultural technology and the interaction between both, e.g. shorter cultivars facilitated higher nitrogen fertilization resulting in higher yields.
Table 1. Acreage, grain yield and total production of bread wheat in the world, in Europe and in Germany; comparison of the years 1971-73, 1991-1993, 2011-2013, and 2018-2020 (3 year means each, FAOSTAT 06/2022)
| Period | Harvest area | Grain yield dt/ha | Production | Production increase vs. 1971-1973 |
World | 2018-2020 | 216.282 | 34.80 | 752.682 | 2.13x |
| 2011-2013 | 218.659 | 31.77 | 694.690 | 1.97x 1.58x |
| 1991-1993 | 222.926 | 25.07 | 558.888 | |
| 1971-1973 | 215.687 | 16.38 | 353.287 |
|
Europe | 2018-2020 | 61.548 | 41.33 | 254.447 | 1.46x |
| 2011-2013 | 57.129 | 37.63 | 215.135 | 1.24x 1.10x |
| 1991-1993 | 62.493 | 30.74 | 190.645 | |
| 1971-1973 | 89.158 | 19.51 | 173.997 |
|
Germany | 2018-2020 | 2.997 | 72.97 | 21.833 | 2.26x |
| 2011-2013 | 3.146 | 74.48 | 23.417 | 2.42x 1.65x |
| 1991-1993 | 2.482 | 64.45 | 15.974 | |
| 1971-1973 | 2.264 | 42.70 | 9.660 |
|
This success story would have been impossible without the findings of Gregor Mendel, who detected and reported patterns of inheritance, which he defined as “heritable characters“. Thinking in factors, resembling structural “genes“, was the basis for a better understanding of the genetic principles of inheritance, and the application of the so-called “Mendel’s Law” in plant breeding has allowed the continuous development of improved cultivars.
Applying these principles, essential traits of crop plants have been continuously improved, e.g. resistance against diseases, pests and product quality. This was possible because these traits are often controlled by relatively few or even single genes. Many of such genes have already been identified decades ago by classical segregation analyses and tests for allelism. For that reason, most of current crop varieties are resistant against major diseases, e.g. wheat and barley against powdery mildew or different rusts. Such “Mendelian genes“ were amongst the first to be investigated by molecular methods and finally isolated combining Mendelian segregation analyses with the findings of Thomas Hunt Morgan (1911 and later) on linkage of genes in genetic mapping and map-based cloning. Molecular markers today facilitate efficient marker-based selection procedures and pyramiding of resistance genes. Furthermore, isolated genes can also be efficiently modified today by genome editing tools such as CRISPR/Cas.
Due to obviously increasing effects of climate change, it will be necessary in the future to develop new varieties with a better tolerance against abiotic stress, e.g. heat, and drought. Traits like yield or yield components are usually not controlled by one or a few genes but several or even many genes. They are determined polygenically, and in addition, these traits are influenced by the interaction of the genotype by the environment. Therefore, no Mendelian classes may be detectable but they show a typical mode of quantitative trait expression i.e. a Gaussian distribution. However, by applying QTL-analyses these traits can be separated into Mendelian factors, so called quantitative trait loci (QTL), i.e. genomic regions explaining a certain amount of the phenotypic variance. This is also true for yield and a number of major quality traits. For breeding and improvement of such complex traits, new approaches – “genomic prediction and selection“ – have been developed recently. Here, genotypes are screened for genetic variation giving hint to a specific, wishful trait expression (phenotype). This way, starting from Mendels laws via the empirical development and application of breeding schemes, a continuous extension and optimization of methodology takes place in todays knowledge-based plant breeding. This will enable the further elevation of yield potential, of yield stability and the quality of crop plant products in the future.
Why did Mendel choose the pea as the experimental plant?
Pea was a common model plant at the time, because keeping of and caring for peas is relatively easy, the plants grow quickly and the generation time is short.
There are easily recognisable characteristics such as plant size, flower colour, pod shape and pod colour, seed shape or cotyledon colour.
The plants are relatively small, so they require little greenhouse or field space.
Peas reproduce by self-fertilisation, so that clear segregation ratios occur in cross progeny.
The plants have a high propagation coefficient, so that large populations are available for genetic analyses.
In this context, it was a fundamental advantage for the experimental results and their genetic interpretation that peas are diploid (2n=2x=14), so that each gene primarily occurs in only two variants (alleles). Therefore, in a cross of homozygous parental lines the typical Mendel segregation ratios can be observed as a result of the mode of action (dominant, intermediate) and re-combination. If Mendel's experimental plants had been polyploid, the derivation of the genetic rules would undoubtedly have been much more difficult due to the multiple alleles. The same holds true if he had chosen an out-breeding species or an apomictic species, as used in that time by many botanists.
Furthermore, it was an enormous advantage that the traits he analysed were monogenic, i.e. inherited by a single gene. Mendel did not yet use the term "gene" – he used the term "hereditary character". Other terms that were coined in the following periods are: cistron, operon, ORF (open reading frame), and others. Today we know not only the location in the genome and the DNA structure of many genes, but also their function; for example, the A gene for flower colour of pea intervenes in anthocyanin biosynthesis: White flowering peas are natural base exchange mutants (Guanin-> Adenin) (Fig. 1). It was not until decades after Mendel that such gene mutations were experimentally triggered, with the help of ionising radiation or mutagenic chemicals and used in plant breeding. More recently, Hellens et al. (2010) have identified the pea gene A as the factor determining anthocyanin pigmentation in pea. The A gene encodes a bHLH transcription factor. The white flowered mutant allele most likely used by Mendel is a simple G to A transition in a splice site that leads to a mis-spliced mRNA with a premature stop codon. Hellens et al. (2010) have identified a second rare mutant allele: The A2 gene encodes a WD40 protein that is part of an evolutionarily conserved regulatory complex and two premature stop codons were identified in white flowering genotypes.
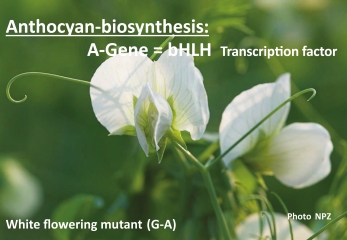

Fig. 1. White-flowered pea plant (Pisum sativum) due to a natural recessive point mutation (Guanin -> Adenin) of the original dominant red wild type. The A-gene represents the transcription factor bHLH intervening in anthocyanin biosynthesis.
In view of the many demands made on our crops in cultivation, processing and use, they must combine a large number of favorable characteristics: starting with tolerance to environmental stress and resistance to pathogens and pests (i.e. yield stability), via yield to product quality, e.g. wheat flour, barley malt, rapeseed cooking oil, plant protein e.g. from legumes.
Knowledge of Mendel's rules has subsequently proved to be extraordinarily fruitful for plant genetics and extremely useful for breeding; in concrete terms, this has resulted in the following consequences, among others: The 1st Mendel rule describes the uniformity of the F1 generation when crossing homozygous parents and is a basis for today's widespread hybrid breeding. The 2nd Mendel rule (segregation rule) states that genetic segregation is to be expected for the first time in the F2 generation, so that the earliest selection possibility exists here and may be conducted for simply inherited traits in line breeding schemes of self-pollinating species. The 3rd Mendel rule (independence rule) states that different traits in the F2 basically split independently of each other; this is the basis for combining favorable genes or alleles for different target traits ("combination or cross-breeding"). However, this holds only true to a limited extend, i.e. when genes are not linked. Linkage between genes was detected by Thomas Hunt Morgan in the 1920s (different references), and is the basis for high density molecular marker maps today available in many crop species (Fig. 2).
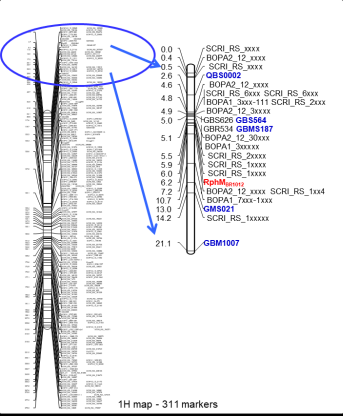
Fig. 2. Dense marker map of barley chromosome 1H comprising the rust resistance locus RphMBR1012 (Figure was taken from Perovic et al., unpublished)
Thus, it was a fortunate circumstance for the further development of genetics that in the case of Mendel's experiments such linkage did not play a role in the traits he described since the causal genes are located on different chromosomes and therefore were independent. Further exceptions to the genetic laws are, for example, parental effects (lack of reciprocity), epistasis (non-allelic gene interaction), polygeny, haploidy or environmental effects (GxE interaction).
If genes are unlinked, relevant alleles, for example a cross between a virus-resistant (see below) but mildew-susceptible parent plant (vvMM) and a virus- susceptible but mildew-resistant parent (VVmm), can result in a double-recessive mildew- and equally virus-resistant progeny (mmvv). So called “combination breeding“ has created considerable improvements in crops over many decades, especially in self-pollinators such as barley and wheat. The crossing of two parents P1 and P2 (sometimes extended by crossing the F1 with a third parent) for the purpose of combining favourable characteristics in each case results in an offspring that becomes increasingly homozygous due to selfing in the subsequent generations. Multi-stage selection is basically possible from F2 onwards. Special methods have been developed for breeding practice, which are conducted to a greater or lesser extent even today depending on the conditions of the material and the preference of the breeder; these include the pedigree method, bulk method, the use of single-seed descents, or the direct production of homozygous offspring via in vitro culture of haploid gametes, preferably immature pollen (“haploid method“). In this respect, it has to be noticed that these doubled haploids in case of a dominant/recessive inheritance do not segregate in a 3:1 manner but a 1:1 frequency as there are no heterozygous genotypes, i.e. homozygous recessive genotypes are more frequent.
Multi-stage line selection based on individual plants or plant progenies (bulks) over several generations is always followed by yield tests, initially at the breeding site, then also multi-locational and finally multi-annual. In principle, self-pollinated varieties are still bred in this way today, for example wheat, barley, oats, pea, etc. Of course, the breeding methodology has been further developed and refined over the decades. Cell and tissue culture techniques are now an integral part of the “tool box” of a modern breeding company: Whether for the efficient multiplication of a genotype by cloning, the rapid and pathogen-free propagation by meristem culture, the acceleration of backcrossing steps by embryo culture, the effective production of species or genus hybrids via sexual crossing and embryo rescue or via asexual crossing by cell fusion, the creation of homozygous plants via the “haploid method”, i.e. in vitro culture of gametes, especially microspores (immature pollen), and others. In recent times, molecular genetics has also increasingly found its way into breeding procedures. This has been made possible by the enormous development of -omics technologies such as genomics, transcriptomics, metabolomics, proteomics, etc. The genomes of practically all agriculturally and horticulturally relevant plant species have now been completely sequenced and are available for plant breeders.
Quite a few relevant traits are monogenically controlled, including many disease resistances. For example, powdery mildew (Blumeria graminis f. sp. tritici) causes serious yield losses in bread wheat (T. aestivum) production. In the course of time numerous genes (Pm genes) and quantitative trait loci (QTL) have been identified distributed over the entire wheat genome mediating resistance against powdery mildew; recently, Kang et al. (2020) have provided a summary of over 200 powdery mildew genes and QTL across all 21 chromosomes of bread wheat.
These monogenic resistances are known for many pathogens in wheat, e.g. stripe rust (Yr-genes), leaf rust (Lr-genes), stem rust (Sr-genes), soil-borne cereal virus (Sbm1) or even against the orange wheat blossom midge (Sm1). However, the introduction of disease resistance in elite winter wheat cultivars is a lengthy process as shown here for the pedigree of wheat varieties combining resistance against powdery mildew i.e. Pm17 (1A/1R translocation) and leaf rust (Lr24) plus Pseudocercosporella herpotrichoides (Table 2, H. Kempf, pers. comm. 2022), which can be abridged today by molecular markers.
Table 2. Partial pedigree of elite wheat varieties derived from “wide crosses“ using unadapted resistance sources of wheat (T. aestivum) (Dr. Hubert Kempf, Secobra)
1984 | Kronjuwel × AMIGO (Pm17 + Lr24) |
1985 | Kronjuwel/(Pm17 + Lr24) × Kronjuwel |
1993 | [Kronjuwel/(Pm17 + Lr24) × Kronjuwel] × Piko |
1999 | (Atlantis/Cardos × [Kronjuwel/(Pm17 + Lr24) × Kronjuwel] × Piko) ->MEMORY |
2008 | “Memory“ (Pm17 + Lr24) × JB Asano -> ASORY |
2013 | Registration MEMORY (Pm17 + Lr24) |
2018 | Registration ASORY (Pm17 + Lr24) |
2019 | Registration CAMPESINO (Pm17 + Lr24 + Pch1) |
Pm = powdery mildew, Lr = leaf rust, Pch = Pseudocercosporella
Another example are the so-called “green revolution genes”. These genes have a major effect on plant height as they cause – depending on the respective allele – dwarf or semi-dwarf growth. An example is shown in Figure 3, which illustrates the effect of different Rht-B1 and Rht-D1 alleles, i.e. genes causing gibberellic (GA)-insensitivity derived from the Japanese wheat variety Norin10. These semi dwarf genotypes are much lesser sensitive to lodging facilitating higher nitrogen fertilization resulting in higher yield.

Fig. 3. Phenotypes of near-isogenic lines (NIL) derived from wheat cv. Mercia, differing in Rht- B1 and Rht-D1 alleles; Rht-1 represents the original variety (Photo: John Innes Centre, Norwich/UK, Tony Worland, see also Li et al., 2006)
Besides, quality characteristics are also controlled by single genes. An illustrative example is oil quality of rapeseed (B. napus) and other oil crops. Unsaturated fatty acids – derived from stearic acid (C18:0), via oleic (C18:1) and linoleic acid (C18:2) to linolenic acid (C18:3) are formed in several steps by desaturase enzymes encoded by structural genes. In addition, long-chain fatty acids are formed by elongation in crucifers (= Brassicaceae) incl. oilseed rape, Brassica napus): C18:1 -> C20:1 (eicosenic) -> C22:1 (erucic acid). The combination of functional alleles and their additive effects enable maximum erucic acid content of so-called High Erucic Acid Rapeseed (HEAR). For details see Figure 4 (Bates et al., 2013).
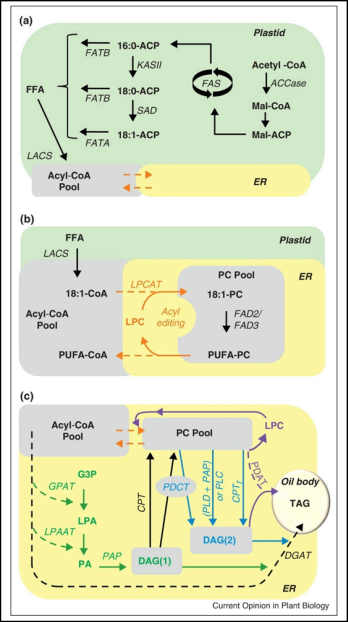
Fig. 4. (a) Plastid fatty acid biosynthesis, (b) Acyl editing and (c) Triacylglyceride (TAG) synthesis. Acyl transfer reactions are dashed lines. Green lines are de novo TAG synthesis, blue lines are PC-derived DAG synthesis, orange lines are acyl editing, and purple represents phospholipid:diacylglycerol acyltransferase (PDAT). DAG(1) is de novo synthesized DAG and DAG(2) is PC-derived DAG. Abbreviations: substrates are in bold: ACP, acyl carrier protein; DAG, diacylglycerol; FFA, free fatty acid; G3P, glycerol-3-phosphate; LPA, lyso-phosphatidic acid; LPC, lyso-phosphatidylcholine; Mal, malonate; PA, phosphatidic acid; PC, phosphatidylcholine; PUFA, polyunsaturated fatty acids; TAG, triacylglycerol. Enzymatic reactions are in italics: ACCase, acetyl-CoA carboxylase; CPT, CDP-choline:DAG cholinephosphotransferase; DGAT, acyl-CoA:DAG acyltransferase; FAD, fatty acid desaturase; FAS, fatty acid synthase; FATA, acyl-ACP thioesterase A; FATB, acyl-ACP thioesterase B; GPAT, acyl-CoA:G3P acyltransferase; KASII, ketoacyl-ACP synthase II; LACS, long chain acyl-CoA synthetase; LPAAT, acyl-CoA:LPA acyltransferase; LPCAT, acyl-CoA:LPC acyltransferase; PAP, PA phosphatase; PDCT, PC:DAG cholinephosphotransferase; PLC, phospholipase C; PLD, phospholipase D; SAD, Stearoyl-ACP desaturase. (Figure was taken from Bates et al., 2013, CC BY-NC-ND 3.0 license)
Barley yellow mosaic virus disease was first detected in Japan in 1940 (Ikata & Kawai, 1940) and is now present in many East Asian countries and especially in Europe (Kühne, 2009). There, barley yellow mosaic virus has become one of the most important diseases of winter barley since its first detection in Germany in 1978 (Kühne, 2009) leading to severe damage and yield losses of up to 50% (see Fig. 5).

Fig. 5. Field-grown winter barley plants: barley yellow mosaic virus resistant variety (right) in comparison to a virus-sensitive variety (Northern Germany).
Both, Barley yellow mosaic virus (BaYMV) and Barley mild mosaic virus (BaMMV) from which different strains are known (Habekuss et al., 2008) are bymoviruses, which are transmitted by the soil-borne plasmodiophorid Polymyxa graminis (Adams & Swaby, 1988). Resistance to BaMMV/BaYMV has been detected in Germany soon after the discovery of these viruses within the set of released cultivars, and it was shown by classical segregation analysis using BaMMV, which in contrast to BaYMV is easily mechanical transmissible (Friedt, 1983), that resistance is due to a single recessive gene, later on called rym4 (Friedt & Foroughi-Wehr, 1985, Friedt et al., 1989). Extensive screening programs explained that resistance to Barley yellow mosaic virus is quite frequent within the barley primary gene pool (e.g. Ordon & Friedt, 1993; Habekuss et al., 2008). Mendelian segregation analyses of crosses of these resistant genotypes to carriers of rym4, it turned out that recessive resistance genes different from rym4 are present demonstrated by a good fit to a segregation ratio of 7r:9s indicative for two independent recessive resistance genes causing resistance each (Götz & Friedt, 1993). In a next step, by intercossing donors of resistance carrying genes different from rym4 it was shown that these carry different recessive resistance genes (Ordon & Friedt, 1993). In addition, two dominant resistance genes have been identified in the secondary gene pool, i.e. Hordeum bulbosum (Ruge et al., 2003; Ruge-Wehling et al., 2006), and a first dominant resistance gene has been identified meanwhile also in cultivated barley (Kai et al., 2012).
These classical Mendelian segregation analyses together with the knowledge of linkage of genes developed by Thomas Hunt Morgan has been the basis for developing molecular markers (Graner & Bauer, 1993) mainly based on the analyses of DH-lines allowing a replicated test for virus resistance in contrast to F2-plants (for overview see Tuvesson et al., 2021).
Based on these markers, different marker-based selection procedures have been developed. Together with doubled haploid techniques, which are routinely used in barley breeding today, respective markers facilitate a reliable and fast selection for virus resistance, e.g. doubled haploid populations may be screened directly in vitro and only those plantlets carrying the resistance encoding allele have to be transferred to the greenhouse. In general, donors of new BaMMV/BaYMV resistance genes are rather un-adapted to productive growing systems (e.g. Ordon & Friedt, 1994). To combine virus resistance with superior agronomic performance, time-consuming backcrossing procedures are needed. This holds especially true when recessive resistance genes have to be incorporated like in the case of BaMMV/BaYMV, just as a selfing generation is needed after each backcross to identify homozygous recessive genotypes on the phenotypic level (3s:1r). In contrast to this, in marker-based procedures, the recessive resistance encoding allele can be directly followed by a co-dominant marker or a dominant one showing an additional fragment linked to the resistance encoding allele (Ordon et al., 2003; 2009), thus saving one generation per backcrossing cycle. Furthermore, molecular markers facilitate efficient pyramiding of resistance genes especially in combination with doubled haploids (Werner et al., 2005; 2007), thereby prolonging the use of partly overcome resistance genes.
Besides the direct use in breeding, Mendelian segregation and knowledge on linkage has also been the basis for isolating resistance genes rym4/5 and rym11 via a map based cloning approach (Stein et al., 2005; Yang et al., 2014) which is based on constructing a high resolution mapping population followed by marker saturation, i.e. segregation and linkage. It turned out that the rym4/5 locus comprises the translation initiation factor 4e gene Hv-eIF4E (Stein et al., 2005; Kanyuka et al., 2005) and rym11 a protein disulfide isomerase (HvPDIL5-1). Meanwhile both of these genes have been edited using CRISPR/Cas9 (Hoffie et al., 2021, cf. Fig. 6) and the isolation of different resistance genes e.g. rym15 using recent genomic tools is underway (Wang et al., 2022).
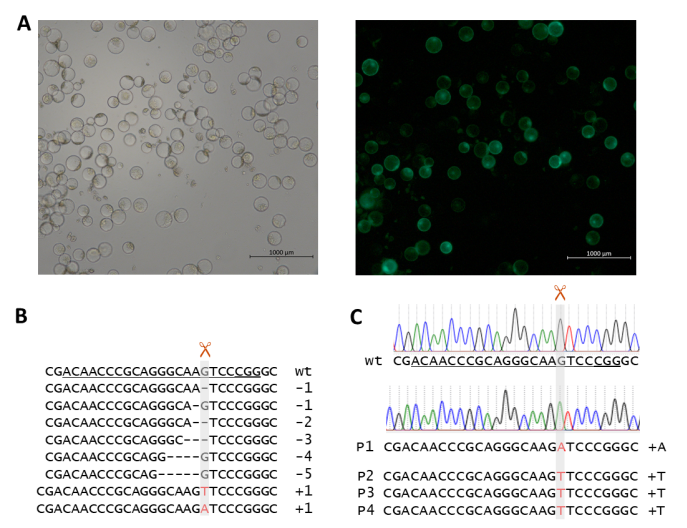
Fig. 6: (A) Mesophyll protoplasts of winter barley cv. “Igri” after PEG-mediated transformation with a GFP-carrying vector, after 60 h of incubation at 21°C. These protoplasts serving as positive control for viability and genetic transformation, respectively, were recorded using bright field (left) and epifluorescence (right) microscopy. (B) Deep-sequencing of amplicons of target region 1 after transformation of barley protoplasts using vectors carrying cas9 and target motif 1-specific gRNA expression units. (C) Chromatograms of Sanger sequencing target region 1 of wild-type “Igri” as compared with primary mutant plants. The unambiguous DNA sequence indicates homozygosity of the inserted A nucleotide. Grey vertical bars: cleavage positions, horizontal lines: deletions, red letters: insertions, wt: wild-type sequence, target motifs underlined with PAMs being double underlined, +A: 1-bp insertion of adenine nucleobase, +T: 1-bp insertion of thymine nucleobase. Results of PEG-mediated transformation of protoplast transformation of barley cv. ‘Igri’ using GFP-carrying vector (Figure was taken from Hoffie et al., 2021, CC-BY 4.0 license)
The story of resistance to BaYMV/BaMMV clearly elucidates the importance of Mendel´s law and its useful application in plant breeding and research. Genome editing has also been applied to many further plant species, e.g. for improving mildew resistance in bread wheat (Li et al., 2022).
Oligo- or polygenic inheritance plays a major role concerning traits of agronomic importance, i.e. quantitative traits. Such traits are not accessible to a classical genetic analysis according to Mendel. Here, however, combined quantitative-genetic and molecular-genetic approaches made it possible to depict associations of special genome regions with respective traits (QTL mapping in biparental populations or via genome wide association studies (GWAS)). For example, Zymoseptoria tritici causing Septoria tritici leaf blotch, is a difficult fungus to control because of its extremely high genetic variability making it one of the most important wheat pathogens today. Recently, Karlstedt et al. (2019) have detected 4 QTL in wheat explaining between 8.5 -17.5% of the phenotypic variance. (Fig. 7). Thus, like Mendel genes, they can now be made accessible for targeted selection with the help of associated genetic markers.
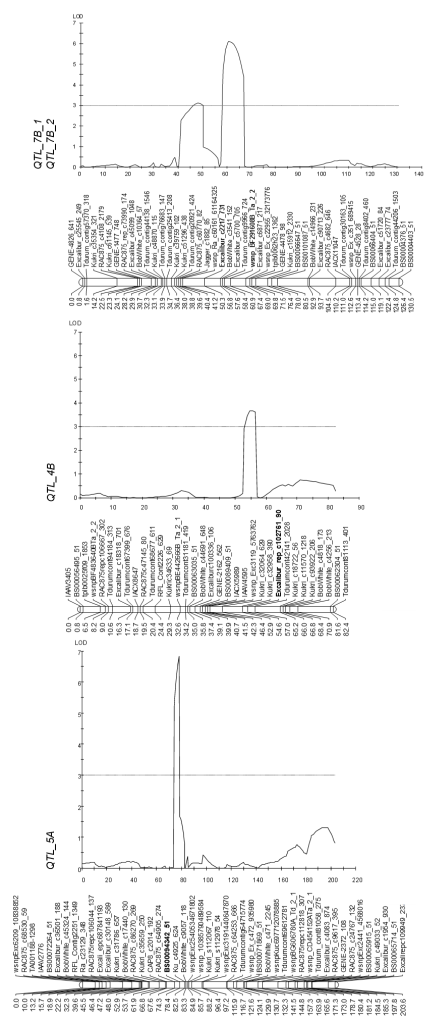
Fig. 7: Results of QTL analyses for Zymoseptoria tritici resistance in the DH population HTRI1410 × susceptible parental lines (Reprinted by permission from Springer Nature: Euphytica: “Mapping of quantitative trait loci (QTL) for resistance against Zymoseptoria tritici in the winter spelt wheat accession HTRI1410 (Triticum aestivum subsp. spelta)”, Karlstedt et al., 2019).
The same applies to the plant response to abiotic stress, such as cold, heat or drought. Resistance or tolerance to such stressors is at best partial and is inherited quantitatively. Here, too, however, it may help to break down the quantitative trait complexes into "Mendelian" units to gain access to individual synthesis steps or metabolic pathways and the genes underlying these (Wehner et al., 2015). Typically, signal transduction chains are effective here: for example, sensor enzymes can activate phytohormones (e.g. abscisic acid, ABA), which in turn activate genetic switches (transcription factors) such as CAMTA, DREB, MYB or bZIP to set defence or repair mechanisms in motion. This brief description shows that numerous genes must be functionally involved and that, due to the complex genetics, the breeding of such traits is difficult and therefore inevitably lengthy, but can be conducted today in a directed manner, by employing molecular markers.
We owe Gregor Mendel a basic understanding of the inheritance of plant characteristics:
Hereditary “factors” (genes) determine the appearance of organisms (phenotype), but can be modified by “environmental effects”.
In diploid individuals, the factors basically occur in two forms (alleles), they can be recombined by crossing and thus lead to new phenotypes with novel trait combinations.
These findings were the basis for the development of breeding methods from line to hybrid breeding, combined with the invention of improved breeding schemes.
Their systematic use in classical plant breeding has led to enormous improvements in crops: today's varieties are generally superior to older varieties in terms of disease resistance, stress tolerance, yield and product quality (flour, edible oil, protein, animal feed, etc.).
The useful addition of biotechnological approaches in the form of cellular and molecular biological work steps is increasingly turning empirical plant breeding into knowledge-based plant breeding.
The authors declare that they do not have any conflicts of interest.
Adams, M.J., A.G. Swaby, 1988: Confirmation of the transmission of barley yellow mosaic virus (BaYMV) by the fungus Polymyxa graminis. Annals of Applied Biology, 112 (1), 133-141, DOI: 10.1111/j.1744-7348.1988.tb02048.x.
Bates P.D., S. Stymne, J. Ohlrogge, 2013: Biochemical pathways in seed oil synthesis. Current Opinion in Plant Biology 16 (3), 358-364, DOI: 10.1016/j.pbi.2013.02.015.
Friedt, W., 1983: Mechanical transmission of soil‐borne barley yellow mosaic virus. Journal of Phytopathology, 106, 16-22, DOI: 10.1111/j.1439-0434.1983.tb00023.x.
Friedt, W., B. Foroughi-Wehr, 1985: Genetics of resistance to barley yellow mosaic virus. Mitteilungen aus der Biologischen Bundesanstalt für Land- und Forstwirtschaft Berlin-Dahlem (Germany).
Friedt, W., R. Götz, R. Kaiser, B. Foroughi-Wehr, 1989: Present state and prospects of breeding for resistance or immunity to barley yellow mosaic virus 1. EPPO Bulletin, 19 (3), 563-571, DOI: 10.1111/j.1365-2338.1989.tb00433.x.
Graner A., E. Bauer, 1993: RFLP mapping of the ym4 virus resistance gene in barley. Theoretical and Applied Genetics 86, 689–693, DOI: 10.1007/BF00222657.
Götz, R., W. Friedt, 1993: Resistance to the barley yellow mosaic virus complex – differential genotypic reactions and genetics of BaMMV‐resistance of barley (Hordeum vulgare L.). Plant Breeding, 111 (2), 125-131, DOI: 10.1111/j.1439-0523.1993.tb00618.x.
Habekuss, A., T. Kühne, I. Krämer, F. Rabenstein, F. Ehrig, B. Ruge‐Wehling, F. Ordon, 2008: Identification of Barley mild mosaic virus isolates in Germany breaking rym5 resistance. Journal of Phytopathology, 156 (1E), 36-41, DOI: 10.1111/j.1439-0434.2007.01324.x.
Hellens, R.P., C. Moreau, K. Lin-Wang, K.E. Schwinn, S.J. Thomson, M.W.E.J. Fiers, T.J. Frew, S.R. Murray, J.M.I. Hofer, J.M. E. Jacobs, K.M. Davies, A.C. Allan, A. Bendahmane, C.J. Coyne, G.M. Timmerman-Vaughan, T.H.N. Ellis, 2010: Identification of Mendel's white flower character. PLoS ONE 5 (10): e13230, DOI: 10.1371/journal.pone.0013230.
Hoffie, R.E., I. Otto, D. Perovic, N. Budhagatapalli, A. Habekuß, F. Ordon, J. Kumlehn, 2021: Targeted knockout of eukaryotic translation initiation factor 4E confers Bymovirus resistance in winter barley. Frontiers in Genome Editing, 3, 784233, DOI: 10.3389/fgeed.2021.784233.
Ikata, S., I. Kawai, 1940: Studies on wheat yellow mosaic disease, No. 154, 1-123. Ministry of Agriculture and Forestry (Japan), URL: https://www.cabdirect.org/cabdirect/abstract/20057005319?q=(do%3a%22Noji-kairyo-shiryo%2c + Ministry + of + Agriculture + and + Forestry.%22).
Kai, H., K. Takata, M. Tsukazaki, M. Furusho, T. Baba, 2012: Molecular mapping of Rym17, a dominant and rym18 a recessive barley yellow mosaic virus (BaYMV) resistance genes derived from Hordeum vulgare L. Theoretical and Applied Genetics, 124 (3), 577-583, DOI: 10.1007/s00122-011-1730-5.
Kang Y., M. Zhou, A. Merry, K. Barry, 2020: Mechanisms of powdery mildew resistance of wheat – a review of molecular breeding. Plant Pathology 69 (4), 601-617, DOI: 10.1111/ppa.13166.
Kanyuka K., A. Druka, D.G. Caldwell, A. Tymon, M. McCallum, R. Waugh, M.J. Adams, 2005: Evidence that the recessive bymovirus resistance locus rym4 in barley corresponds to the eukaryotic translation initiation factor 4E gene. Molecular Plant Pathology 6 (4), 449–458, DOI: 10.1111/j.1364-3703.2005.00294.x.
Karlstedt, F., D. Kopahnke, D. Perovic, A. Jacobi, K. Pillen, F. Ordon, 2019: Mapping of quantitative trait loci (QTL) for resistance against Zymoseptoria tritici in the winter spelt wheat accession HTRI1410 (Triticum aestivum subsp. spelta). Euphytica 215, 108, DOI: 10.1007/s10681-019-2432-3.
Kühne, T., 2009: Soil-borne viruses affecting cereals – Known for long but still a threat. Virus Research, 141 (2), 174-183, DOI: 10.1016/j.virusres.2008.05.019.
Li, S., D. Lin, Y. Zhang, M. Deng, Y. Chen, B. Lv, B. Li, Y. Lei, Y. Wang, L. Zhao, Y. Liang, J. Liu, K. Chen, Z. Liu, J. Xiao, Z. Liu, J.-L. Qiu, C. Gao, 2022: Genome-edited powdery mildew resistance in wheat without growth penalties. Nature 602, 455–460, DOI: 10.1038/s41586-022-04395-9.
Li, X.P., S.Q. Lan, Y.P. Liu, M.D. Gale, T.J. Worland, 2006: Effects of different Rht-B1b, Rht-D1b and Rht-B1c dwarfing genes on agronomic characteristics in wheat. Cereal Research Communications 34, 919–992, DOI: 10.1556/CRC.34.2006.2-3.220.
Morgan, T.H., 1911: Random Segregation versus coupling in Mendelian inheritance. Science 34, 873 p. 384, DOI: 10.1126/science.34.873.384.
Ordon, F., W. Friedt, 1994: Agronomic traits of exotic barley germplasms resistant to soil-borne mosaic-inducing viruses. Genetic Resources and Crop Evolution, 41 (1), 43-46, DOI: 10.1007/BF00051422.
Ordon, F., W. Friedt, 1993: Mode of inheritance and genetic diversity of BaMMV resistance of exotic barley germplasms carrying genes different from ‘ym4’. Theoretical and Applied Genetics, 86 (2), 229-233, DOI: 10.1007/BF00222083.
Ordon, F., K. Werner, B. Pellio, A. Schiemann, W. Friedt, A. Graner, 2003: Molecular breeding for resistance to soil-borne viruses (BaMMV, BaYMV, BaYMV-2) of barley (Hordeum vulgare L.). Journal of Plant Diseases and Protection, 110 (3), 287-295.
Ordon, F., A. Habekuss, U. Kastirr, F. Rabenstein, T. Kühne, 2009: Virus resistance in cereals: sources of resistance, genetics and breeding. Journal of Phytopathology, 157 (9), 535-545, DOI: 10.1111/j.1439-0434.2009.01540.x.
Ruge, B., A. Linz, R. Pickering, G. Proeseler, P. Greif, P. Wehling, 2003: Mapping of Rym14 Hb, a gene introgressed from Hordeum bulbosum and conferring resistance to BaMMV and BaYMV in barley. Theoretical and Applied Genetics, 107 (6), 965-971, DOI: 10.1007/s00122-003-1339-4.
Ruge-Wehling, B., A. Linz, A. Habekuß, P. Wehling, 2006: Mapping of Rym16 Hb, the second soil-borne virus-resistance gene introgressed from Hordeum bulbosum. Theoretical and Applied Genetics, 113 (5), 867-873, DOI: 10.1007/s00122-006-0345-8.
Stein, N., D. Perovic, J. Kumlehn, B. Pellio, S. Stracke, S. Streng, F. Ordon, A. Graner, 2005: The eukaryotic translation initiation factor 4E confers multiallelic recessive Bymovirus resistance in Hordeum vulgare (L.). Plant Journal 42, 912-922, DOI: 10.1111/j.1365-313X.2005.02424.x.
Tuvesson, S.D., C.T. Larsson, F. Ordon, 2021: Use of Molecular Markers for Doubled Haploid Technology: From Academia to Plant Breeding Companies. In: Segui-Simarro, J.M. (Eds.) Doubled Haploid Technology, pp. 49-72, DOI: 10.1007/978-1-0716-1335-1_3.
Wang, Y., A. Habekuss, M. Jayakodi, M. Mascher, R.J. Snowdon, A. Stahl, J. Fuss, F. Ordon, D. Perovic, 2022: High-Resolution mapping of barley mild mosaic virus resistance gene rym15. Frontiers in Plant Science, 1740, DOI: 10.3389/flps.2022.908170.
Werner, K., W. Friedt, F. Ordon, F., 2005: Strategies for pyramiding resistance genes against the barley yellow mosaic virus complex (BaMMV, BaYMV, BaYMV-2). Molecular Breeding, 16 (1), 45-55, DOI: 10.1007/s11032-005-3445-2.
Werner K, W. Friedt, F. Ordon, 2007: Localisation and combination of resistance genes against soil-borne viruses of barley (BaMMV, BaYMV) using doubled haploids and molecular markers. Euphytica 158, 323–329, DOI: 10.1007/s10681-006-9206-4.
Yang, P., A. Habekuß, F. Ordon, N. Stein, 2014: Analysis of bymovirus resistance genes on proximal barley chromosome 4HL provides the basis for precision breeding for BaMMV/BaYMV resistance. Theoretical and Applied Genetics, 127 (7), 1625-1634, DOI: 10.1007/s00122-014-2324-9.